| Accession | GMS-64-82
URL copied to the clipboard |
| Title | Transcriptome and germ-cell development |
| Submit date | 2025-03-25 14:49:44 |
| Last update date | 2025-03-31 10:39:31 |
| Contact | Sung-Joon PARK park AT hgc.jp The University of Tokyo |
| Sharing | |
| Total file size | 65.63 GB |
| Keywords | Spermatogenesis Development Primordial germ cell |
| Experiment type | Bulk transcriptome analysis |
| Summary | Illumina SRA files were prepared from NCBI GEO; Primordial germ cells (PGC) at embryonic day E13.5 (GSM1064457), E14.5 (ERP015330), E16.5 (ERP015330), E18.5 (ERP015330), postnatal P0.5 (DRP002386), P2 (ERP015330), P7.5 (DRP002386), Spermatogonia; SG (GSM1069640), Spermatocytes; SC (GSM1069641), Spermatids; SD (GSM1069642), Spermatozoa; SZ (GSM1069643), Mouse embryonic fibroblast; MEF (GSM1643254), and Sertoli; SL (GSM1069639) Data for SG, SC, SD, SZ, SL, MEF is available at https://gmsuite.hgc.jp/gms_viewer.php?ACC=GMS-64-83 |
| Citation(s) | https://pubmed.ncbi.nlm.nih.gov/23399596/ |
| Organism | Mouse |
| Cell (Tissue) | primordial germ cell |
| Protocol | |
| Data processing | Step1 (Quality control): Trimmomatic (ver. 0.36) assessed all the FASTQ files by using options "ILLUMINACLIP:adapter_file:2:30:10 LEADING:20 TRAILING:20 MINLEN:3" Step2 (Mapping): HISTA2 (ver.2.0.5) coupled with Bowtie2 (ver. 2.2.9) mapped the reads of each replicate to the mouse reference genome mm10 with options "-k 1 --dta --dta-cufflinks". |
| [691] | E13.5 replicate 1 Browse Browse (All) Mouse (mm10); RNA-Seq |
BAM E13.5_1.bam (2.06 GB) |
| [692] | E13.5 replicate 2 Browse Mouse (mm10); RNA-Seq |
BAM E13.5_2.bam (3.05 GB) |
| [693] | E13.5 replicate 3 Browse Mouse (mm10); RNA-Seq |
BAM E13.5_3.bam (2.33 GB) |
| [694] | E14.5 replicate 1 Browse Mouse (mm10); RNA-Seq |
BAM E14.5_1.bam (2.16 GB) |
| [695] | E14.5 replicate 2 Browse Mouse (mm10); RNA-Seq |
BAM E14.5_2.bam (3.81 GB) |
| [696] | E14.5 replicate 3 Browse Mouse (mm10); RNA-Seq |
BAM E14.5_3.bam (4.31 GB) |
| [702] | E16.5 replicate 10 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_10.bam (4.15 GB) |
| [697] | E16.5 replicate 5 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_5.bam (2.94 GB) |
| [698] | E16.5 replicate 6 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_6.bam (3.70 GB) |
| [699] | E16.5 replicate 7 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_7.bam (3.82 GB) |
| [700] | E16.5 replicate 8 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_8.bam (4.27 GB) |
| [701] | E16.5 replicate 9 Browse Mouse (mm10); RNA-Seq |
BAM E16.5_9.bam (2.66 GB) |
| [703] | E18.5 replicate 1 Browse Mouse (mm10); RNA-Seq |
BAM E18.5_1.bam (1.40 GB) |
| [704] | E18.5 replicate 2 Browse Mouse (mm10); RNA-Seq |
BAM E18.5_2.bam (1.33 GB) |
| [705] | P0.5 replicate 1 Browse Mouse (mm10); RNA-Seq |
BAM P0.5_1.bam (3.18 GB) |
| [706] | P0.5 replicate 2 Browse Mouse (mm10); RNA-Seq |
BAM P0.5_2.bam (5.49 GB) |
| [707] | P2.0 replicate 3 Browse Mouse (mm10); RNA-Seq |
BAM P2.0_3.bam (1.54 GB) |
| [708] | P2.0 replicate 4 Browse Mouse (mm10); RNA-Seq |
BAM P2.0_4.bam (1.70 GB) |
| [709] | P2.0 replicate 5 Browse Mouse (mm10); RNA-Seq |
BAM P2.0_5.bam (1.25 GB) |
| [710] | P2.0 replicate 6 Browse Mouse (mm10); RNA-Seq |
BAM P2.0_6.bam (1.27 GB) |
| [711] | P7.5 replicate 1 Browse Mouse (mm10); RNA-Seq |
BAM P7.5_1.bam (3.18 GB) |
| [712] | P7.5 replicate 2 Browse Mouse (mm10); RNA-Seq |
BAM P7.5_2.bam (6.02 GB) |
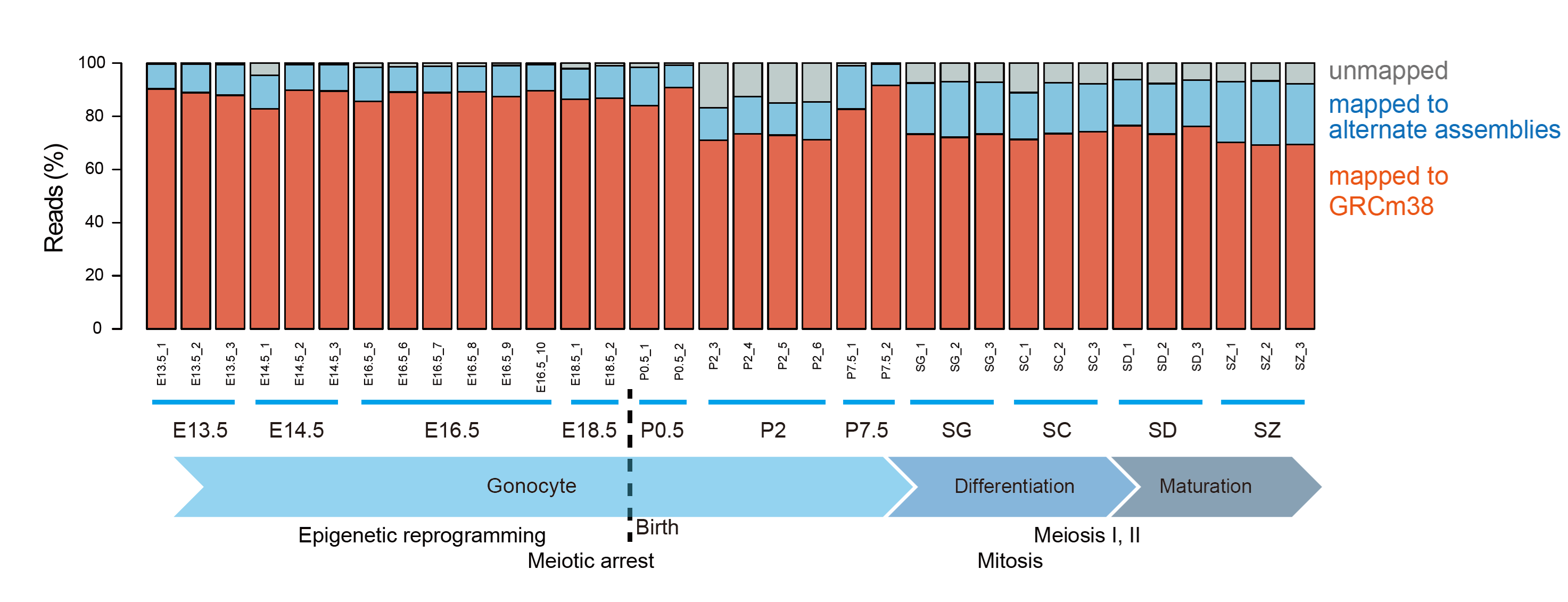
| [721] | RNA-seq mappability in germ-cell development germcell_dev.png (177.55 KB) |

|
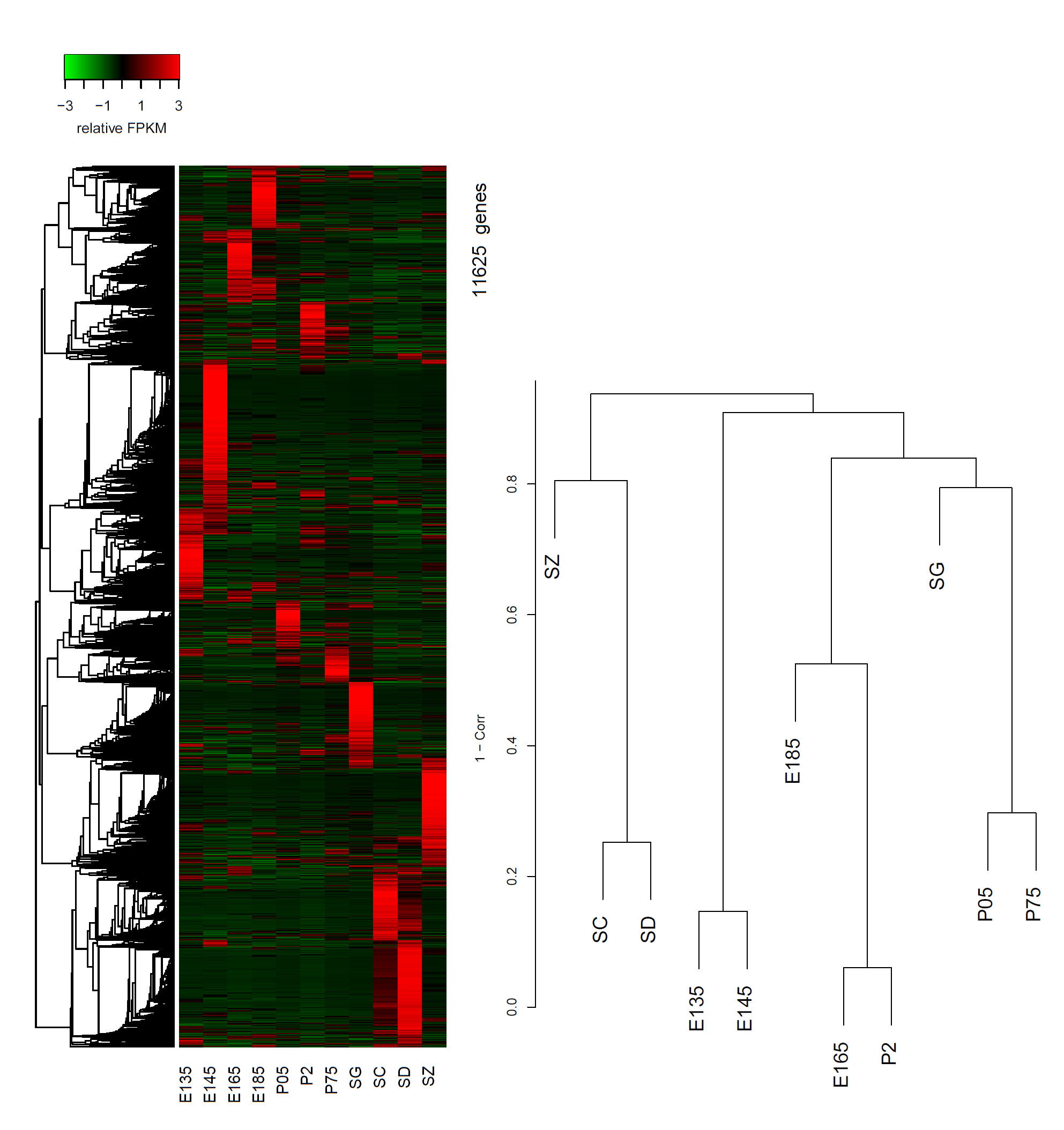
| [735] | Gene expression clustering transcriptome_changes.png (187.43 KB) |
|
| [761] | Gene expression profile (TPMs from E13.5 to SZ with SL and MEF) |
expression_profile.txt (2.31 MB) |
| [691] E13.5_1.bam
E13.5 replicate 1 application/octet-stream 2.06 GB MD5: 43c96865342845684a2c024bd5fb7c88 |
| [692] E13.5_2.bam
E13.5 replicate 2 application/octet-stream 3.05 GB MD5: e5c97aa89b273afd4f5a81047dff46c1 |
| [693] E13.5_3.bam
E13.5 replicate 3 application/octet-stream 2.33 GB MD5: aeb0da532ca1be6c40035c3b53b5d4d4 |
| [694] E14.5_1.bam
E14.5 replicate 1 application/octet-stream 2.16 GB MD5: 8b336d04f51195acacb43a5de9a32ed8 |
| [695] E14.5_2.bam
E14.5 replicate 2 application/octet-stream 3.81 GB MD5: d4029813594961d7c3d66e7cd6002274 |
| [696] E14.5_3.bam
E14.5 replicate 3 application/octet-stream 4.31 GB MD5: c5546d9b0d77c65e3c4076594650d855 |
| [702] E16.5_10.bam
E16.5 replicate 10 application/octet-stream 4.15 GB MD5: 350994d33416dbc3dc8a813811298339 |
| [697] E16.5_5.bam
E16.5 replicate 5 application/octet-stream 2.94 GB MD5: 3781b6b928b6a9629c979444118298ef |
| [698] E16.5_6.bam
E16.5 replicate 6 application/octet-stream 3.70 GB MD5: 35188fc35e8db2dc50a6502abc97f98a |
| [699] E16.5_7.bam
E16.5 replicate 7 application/octet-stream 3.82 GB MD5: 4512455a8dcc990172266a0c4417129e |
| [700] E16.5_8.bam
E16.5 replicate 8 application/octet-stream 4.27 GB MD5: ba50ce2d59a753154d3ce0772ffc3659 |
| [701] E16.5_9.bam
E16.5 replicate 9 application/octet-stream 2.66 GB MD5: 5828374178ccbd1d831937f1babe7387 |
| [703] E18.5_1.bam
E18.5 replicate 1 application/octet-stream 1.40 GB MD5: 913136a4158fdb547809714d55332bf6 |
| [704] E18.5_2.bam
E18.5 replicate 2 application/octet-stream 1.33 GB MD5: 8c084d2a906901423d6cc7c892c671fc |
| [705] P0.5_1.bam
P0.5 replicate 1 application/octet-stream 3.18 GB MD5: 3a2413f3987587020e5f5687ff13937a |
| [706] P0.5_2.bam
P0.5 replicate 2 application/octet-stream 5.49 GB MD5: aac692314a8fc919883818af94ad1754 |
| [707] P2.0_3.bam
P2.0 replicate 3 application/octet-stream 1.54 GB MD5: 82d91053466bc65a71e57d421d4ee8d4 |
| [708] P2.0_4.bam
P2.0 replicate 4 application/octet-stream 1.70 GB MD5: 2ae00a5dc650373b016f6aa5ae237244 |
| [709] P2.0_5.bam
P2.0 replicate 5 application/octet-stream 1.25 GB MD5: 1d44f2f4261cdec72b81d454311debca |
| [710] P2.0_6.bam
P2.0 replicate 6 application/octet-stream 1.27 GB MD5: 90caa208294ac8a48422d0687634e6eb |
| [711] P7.5_1.bam
P7.5 replicate 1 application/octet-stream 3.18 GB MD5: 3fc7e7859b8b78e325a4339b53cab9ab |
| [712] P7.5_2.bam
P7.5 replicate 2 application/octet-stream 6.02 GB MD5: ae7e1033c176e4bbbbcfc28adafa1b9b |
| [721] germcell_dev.png
RNA-seq mappability in germ-cell development image/png 177.55 KB MD5: 3e720e1dd48b028300e4f23dfac85236 |
| [735] transcriptome_changes.png
Gene expression clustering image/png 187.43 KB MD5: 351cfe17b8e534680d9150794169b392 |
| [761] expression_profile.txt
Gene expression profile (TPMs from E13.5 to SZ with SL and MEF) text/plain 2.31 MB MD5: ff9fd822a15c48f435d60721ff214e93 |