| Accession | GMS-65-112
URL copied to the clipboard |
| Title | A common molecular mechanism underlying Cornelia de Lange and CHOPS syndromes |
| Submit date | 2025-04-01 18:26:35 |
| Last update date | 2025-05-08 20:14:02 |
| Contact | GMSuite GENOMEMODALITY gmsuite AT hgc.jp The University of Tokyo |
| Sharing | |
| Total file size | 2.40 GB |
| Keywords | Disease CHOPS CdLS Nuclear-scale AFF4 NELFC RAD21 NIPBL BRD4 Genome DNA |
| Experiment type | Genome binding/occupancy profiling by high throughput sequencing |
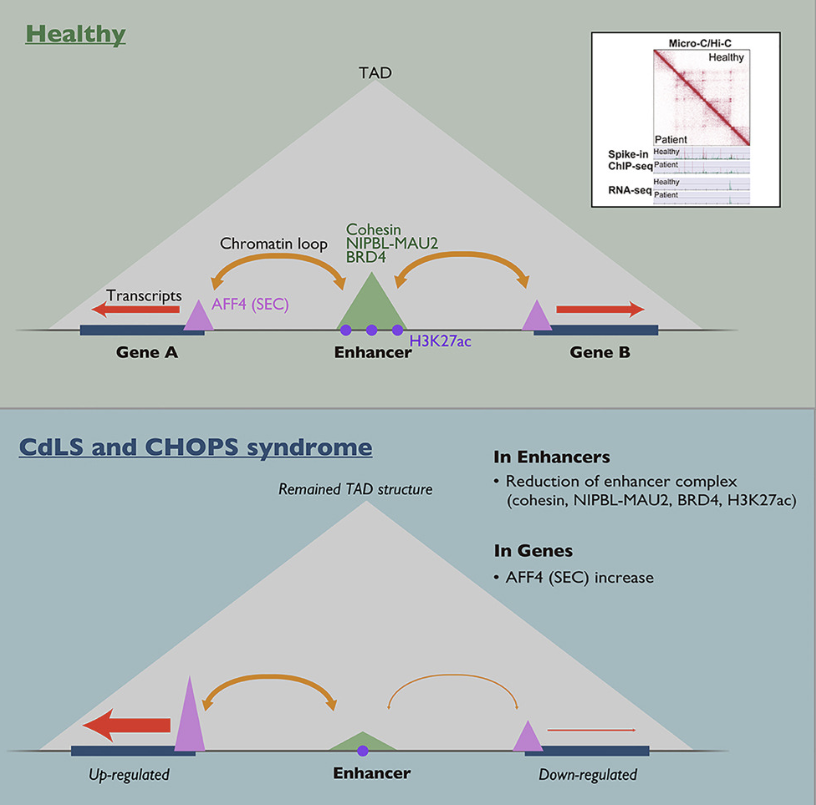
| Summary | For the loading onto chromosomes, cohesin requires the cohesin loader complex formed by NIPBL and MAU2. Cornelia de Lange syndrome (CdLS) is a rare, genetically heterogeneous disorder affecting multiple organs and systems during development, caused by mutations in NIPBL gene, as well as in genes encoding cohesin, a chromatin regulator, BRD4, and cohesin-related factors. CHOPS syndrome that phenotypically overlaps with CdLS is caused by gene mutations of a super elongation complex (SEC) core component, AFF4. In both patient cells, we found a decrease in cohesin, NIPBL, BRD4, and H3K27ac in most enhancers with enhancer-promoter loop attenuation. In contrast, TADs were maintained in both patient cells. |
| Citation(s) | https://pubmed.ncbi.nlm.nih.gov/39983729/ https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE277780 |
| Organism | Human |
| Cell (Tissue) | 293FT |
| Protocol | Chromatin immunoprecipitation was performed as detailed in Nakato et al. (Nature Communications (2023)14:5647). Briefly, human culture cells were crosslinked with 1 % formaldehyde for 10 minutes at room temperature. Chromatin was then sheared by sonication (Branson Sonifier 250D) and combined with mouse spike-in chromatin. The immunoprecipitation was performed overnight at 4°C with an antibody conjugated to Dynabeads Protein A or B (Thermo Fisher Scientific, 10002D or 10004D). After completing the immunoprecipitation and reversing crosslinks, the DNA was purified. Sequencing library was prepared for sequencing using the NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, E7645) according to manufacturer's instructions. |
| Data processing | Sequenced reads were aligned to hg38 and mm10 using Bowtie version 1.2.2 with the options “-n2 -m1”. The uniquely mapped reads were used for subsequent analysis. BigWig files (100-bp bin) were generated by DROMPA version 3.7.2. The read distribution is normalized based on the number of reads derived from the mouse spike-in. BigWig files (100-bp bin) of the normalized read distribution. |
| [745] | RAD21 in wild-type cells Browse Browse (All) Human (hg38); ChIP-Seq |
BW GSM8529572_RAD21_ChIP_FT_control.100.bigwig (151.95 MB) |
| [744] | RAD21 in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529573_RAD21_ChIP_NIPBLmut.100.bigwig (151.69 MB) |
| [746] | NIPBL in wild-type cells Browse Human (hg38); ChIP-Seq |
BW GSM8529574_NIPBL_ChIP_FT_control.100.bigwig (153.77 MB) |
| [747] | NIPBL in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529575_NIPBL_ChIP_NIPBLmut.100.bigwig (153.26 MB) |
| [748] | BRD4 in wild-type cells Browse Human (hg38); ChIP-Seq |
BW GSM8529576_BRD4_ChIP_FT_control.100.bigwig (149.17 MB) |
| [749] | BRD4 in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529577_BRD4_ChIP_NIPBLmut.100.bigwig (148.86 MB) |
| [750] | H3K27ac in wild-type cells Browse Human (hg38); ChIP-Seq |
BW GSM8529578_H3K27ac_ChIP_FT_control.100.bigwig (137.23 MB) |
| [751] | H3K27ac in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529579_H3K27ac_ChIP_NIPBLmut.100.bigwig (135.98 MB) |
| [756] | Input in wild-type cells (replicate 1) Browse Human (hg38); ChIP-Seq |
BW GSM8529580_input_1_FT_control.100.bigwig (153.15 MB) |
| [757] | Input in NIPBL-mutated cells (replicate 1) Browse Human (hg38); ChIP-Seq |
BW GSM8529581_input_1_NIPBLmut.100.bigwig (148.66 MB) |
| [754] | AFF4 in wild-type cells Browse Human (hg38); ChIP-Seq |
BW GSM8529582_AFF4_ChIP_FT_control.100.bigwig (157.90 MB) |
| [755] | AFF4 in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529583_AFF4_ChIP_NIPBLmut.100.bigwig (154.75 MB) |
| [752] | NELFC in wild-type cells Browse Human (hg38); ChIP-Seq |
BW GSM8529584_NELFC_ChIP_FT_control.100.bigwig (153.47 MB) |
| [753] | NELFC in NIPBL-mutated cells Browse Human (hg38); ChIP-Seq |
BW GSM8529585_NELFC_ChIP_NIPBLmut.100.bigwig (149.35 MB) |
| [758] | Input in wild-type cells (replicate 2) Browse Human (hg38); ChIP-Seq |
BW GSM8529586_input_2_FT_control.100.bigwig (150.44 MB) |
| [759] | Input in NIPBL-mutated cells (replicate 2) Browse Human (hg38); ChIP-Seq |
BW GSM8529587_input_2_NIPBLmut.100.bigwig (148.85 MB) |
| [760] | Graphical Abstract GraphicalAbstract.png (303.18 KB) |

|
| [745] GSM8529572_RAD21_ChIP_FT_control.100.bigwig
RAD21 in wild-type cells application/octet-stream 151.95 MB MD5: ca1759b7ef4688fcc75107ac33d40048 |
| [744] GSM8529573_RAD21_ChIP_NIPBLmut.100.bigwig
RAD21 in NIPBL-mutated cells application/octet-stream 151.69 MB MD5: d91f601e3ba049a40219d44c7f1d7cb0 |
| [746] GSM8529574_NIPBL_ChIP_FT_control.100.bigwig
NIPBL in wild-type cells application/octet-stream 153.77 MB MD5: 76e6fc6de5bb062526bd33a18224d8b0 |
| [747] GSM8529575_NIPBL_ChIP_NIPBLmut.100.bigwig
NIPBL in NIPBL-mutated cells application/octet-stream 153.26 MB MD5: 5d2e2e53b58d84b2d248b047194d1865 |
| [748] GSM8529576_BRD4_ChIP_FT_control.100.bigwig
BRD4 in wild-type cells application/octet-stream 149.17 MB MD5: d8c48ff0dc25c3e297f4864ac4ea15db |
| [749] GSM8529577_BRD4_ChIP_NIPBLmut.100.bigwig
BRD4 in NIPBL-mutated cells application/octet-stream 148.86 MB MD5: 6c777ae6d1415e96225b6f5c9c3a0216 |
| [750] GSM8529578_H3K27ac_ChIP_FT_control.100.bigwig
H3K27ac in wild-type cells application/octet-stream 137.23 MB MD5: ef2f498cf74531e4c77a552becc8fc56 |
| [751] GSM8529579_H3K27ac_ChIP_NIPBLmut.100.bigwig
H3K27ac in NIPBL-mutated cells application/octet-stream 135.98 MB MD5: 2aaa5c021f2f8feac1fe204e1b31a01a |
| [756] GSM8529580_input_1_FT_control.100.bigwig
Input in wild-type cells (replicate 1) application/octet-stream 153.15 MB MD5: af93540b592ab49ec7b0b61770b638b7 |
| [757] GSM8529581_input_1_NIPBLmut.100.bigwig
Input in NIPBL-mutated cells (replicate 1) application/octet-stream 148.66 MB MD5: f8ae0408a1ff2d06b4be0e320a71882b |
| [754] GSM8529582_AFF4_ChIP_FT_control.100.bigwig
AFF4 in wild-type cells application/octet-stream 157.90 MB MD5: 1d1a5539b16c25447357a48083fd6724 |
| [755] GSM8529583_AFF4_ChIP_NIPBLmut.100.bigwig
AFF4 in NIPBL-mutated cells application/octet-stream 154.75 MB MD5: 49a83f1bdf4bb0d5f2149d5a9a788924 |
| [752] GSM8529584_NELFC_ChIP_FT_control.100.bigwig
NELFC in wild-type cells application/octet-stream 153.47 MB MD5: 765bcf9279e06eab5f6169aa5898f74f |
| [753] GSM8529585_NELFC_ChIP_NIPBLmut.100.bigwig
NELFC in NIPBL-mutated cells application/octet-stream 149.35 MB MD5: 3b5e4b802a72830109ee24183cc7fe58 |
| [758] GSM8529586_input_2_FT_control.100.bigwig
Input in wild-type cells (replicate 2) application/octet-stream 150.44 MB MD5: 21a961b643f5316fd525837310501d2b |
| [759] GSM8529587_input_2_NIPBLmut.100.bigwig
Input in NIPBL-mutated cells (replicate 2) application/octet-stream 148.85 MB MD5: b25d061388d56efb9f6a604ad54bf66e |
| [760] GraphicalAbstract.png
Graphical Abstract image/png 303.18 KB MD5: 427c61b498e071d6be8da637827ca2e9 |