| Accession | GMS-65-1 (FID: 228)Browse (ALL) |
| Title | Intra-chromosomal Hi-C contacts (mESC Rep1) |
| HiC file | mESC1.intra (1.69 GB) hic mCool Cooler-Tree |
| Submit date | 2022-01-31 12:21:48 |
| Last update date | 2025-05-17 10:45:33 |
| Contact | GMSuite GENOMEMODALITY gmsuite AT hgc.jp The University of Tokyo |
| Sharing | open access |
| Keywords | Chromatin Hi-C PHi-C2 Genome DNA Nuclear-scale |
| Submit date | 2025-01-11 13:01:44 |
| Reference | mm10 (Mouse)
|
| Result file | DomainCall.tar.gz (396.52 MB) |
| Insulation Score (4DN) | 10000bin_100000win.bw 5000bin_100000win.bw |
| GMS Domain | 8164 domains 13683 boundaries |
| GMS Loop | 32989 loops Browse |
| CTCF(+) RAD21(-) | 381 (2.78%) | ||||||||||||||||||||||||||||||||||||||||||||
| CTCF(-) RAD21(+) | 323 (2.36%) | ||||||||||||||||||||||||||||||||||||||||||||
| CTCF(+) RAD21(+) | 2387 (17.45%) | ||||||||||||||||||||||||||||||||||||||||||||
|
GimmeMotifs (Pvalue < 0.001, Top10) |
|
||||||||||||||||||||||||||||||||||||||||||||
|
CTCF&RAD21 in Boundaries Raw ROC file Enriched_motifs.html All_motifs.html |
| CTCF(+) RAD21(-) | 688 (2.09%) LA 736 (2.23%) RA 17 (0.05%) LA & RA |
||||||||||||||||||||||||||||||||||||||||||||
| CTCF(-) RAD21(+) | 785 (2.38%) LA 801 (2.43%) RA 32 (0.1%) LA & RA |
||||||||||||||||||||||||||||||||||||||||||||
| CTCF(+) RAD21(+) | 5050 (15.31%) LA 5169 (15.67%) RA 1261 (3.82%) LA + RA |
||||||||||||||||||||||||||||||||||||||||||||
| CTCF directionality (33005 cases) |
848 (2.57%)Convergent ≫ ≪ 164 (0.5%)Divergent ≪ ≫ 581 (1.76%)Tandem ≫ ≫ 8489 (25.72%)Singleton ≫ |
||||||||||||||||||||||||||||||||||||||||||||
|
GimmeMotifs (Pvalue < 0.001, Top10) LA+RA |
|
||||||||||||||||||||||||||||||||||||||||||||
|
CTCF&RAD21 in Loop Anchors CTCF directionality Raw ROC file Enriched_motifs.html All_motifs.html |
%>4DN_get_insulation_scores_and_boundaries --bweak 0.2 --bstrong 0.5 --cutoff 2 --pixels_frac 0.66
"--binsize 5000 --window 100000,--binsize 10000 --window 100000"
%>java -Xmx32g -jar juicer_tools.jar arrowhead --threads 4 -k KR -m 2000
%>java -Xmx32g -jar juicer_tools.jar hiccups --cpu --threads 4 -k KR -m 1024 -f .1,.1,.1 -p 4,2,1 -i 7,5,3 -t 0.02,1.5,1.75 -d 20000,20000,50000 -r 5000,10000,25000
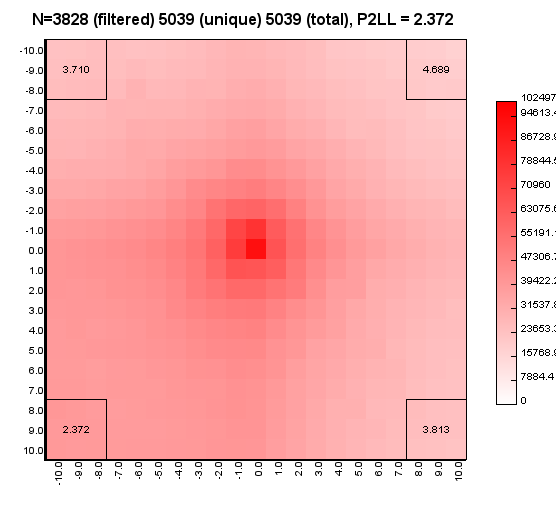
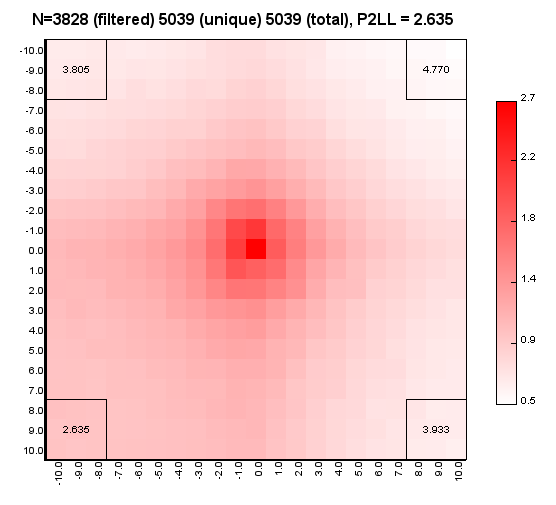
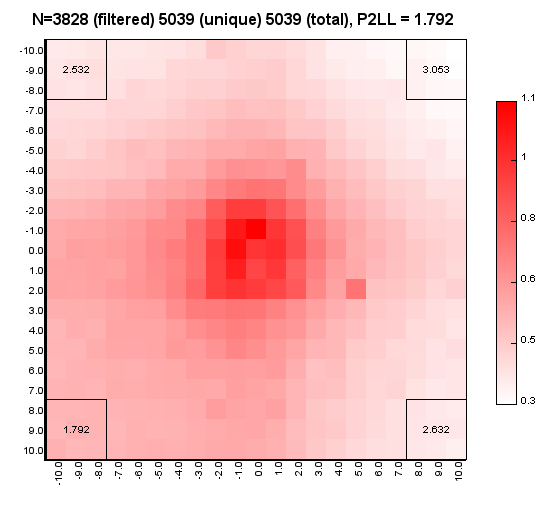
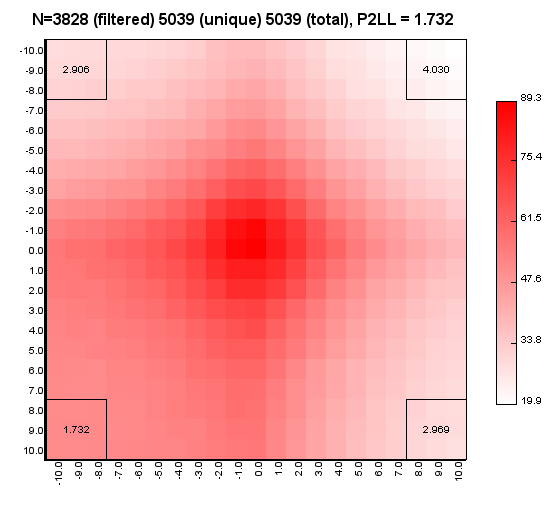
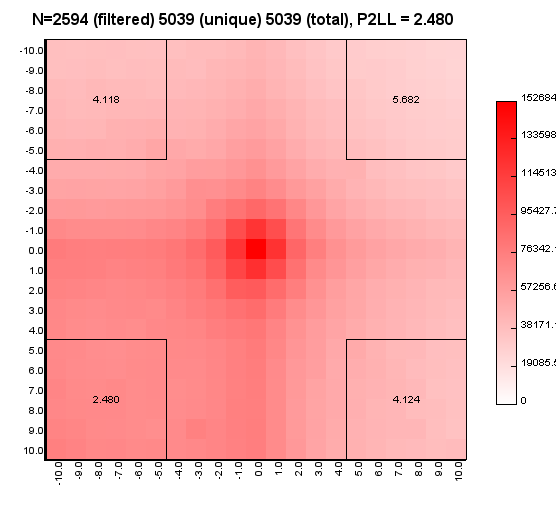
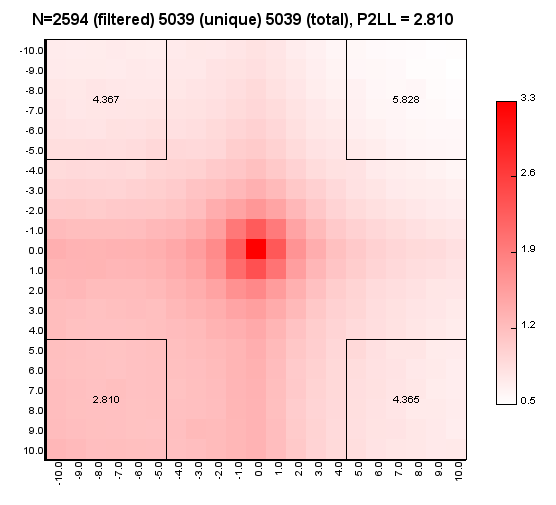
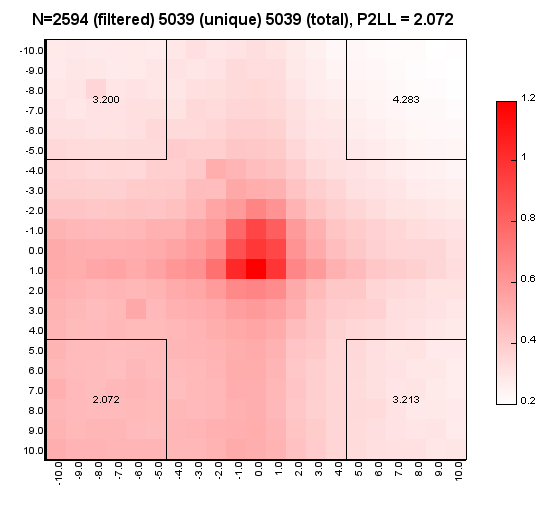
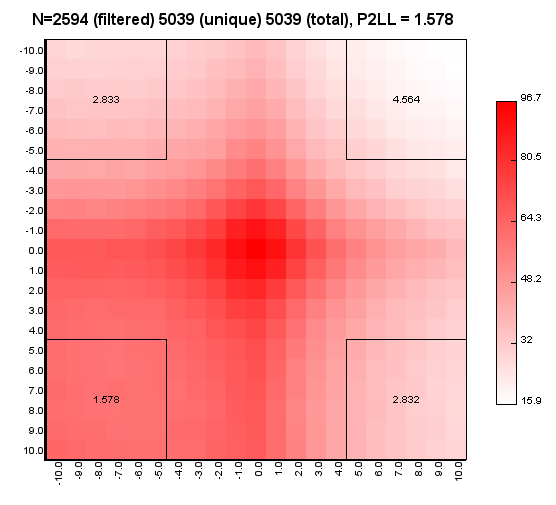
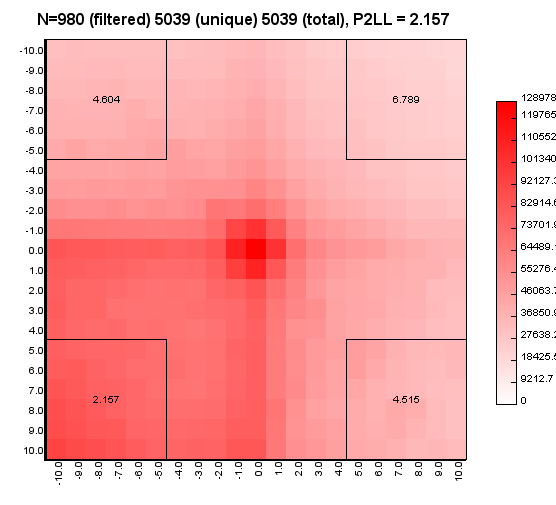
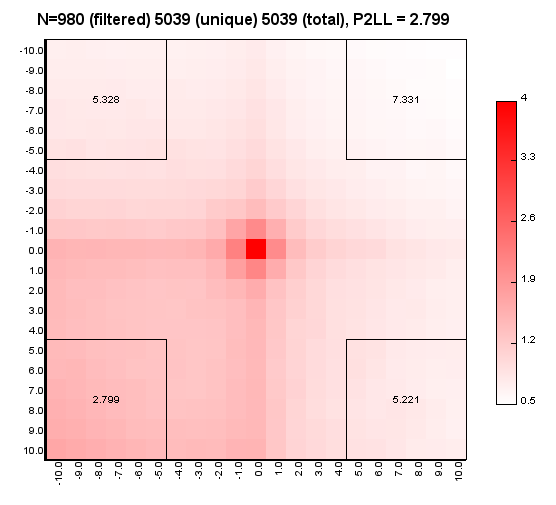
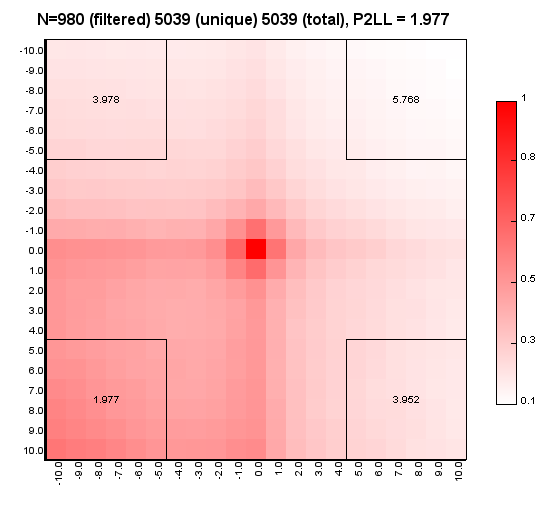
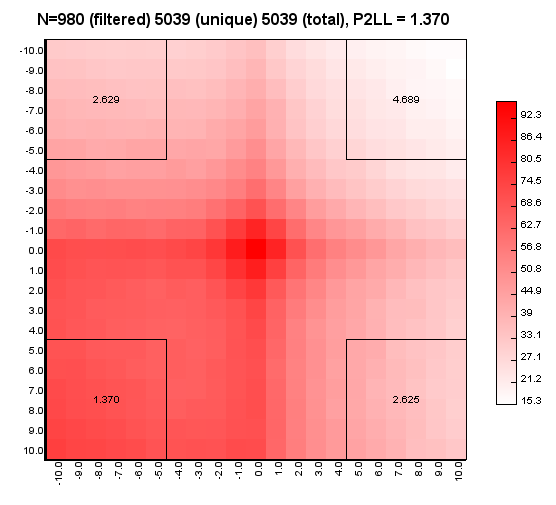
%>java -Xmx32g -jar juicer_tools.jar apa --threads 4
%>hicDetectLoops --threads 2 --peakWidth 2 --windowSize 5 --pValuePreselection 0.1 --peakInteractionsThreshold 10 --obsExpThreshold 1.5 --pValue 0.025 --maxLoopDistance 2000000 --threadsPerChromosome 1 --expected mean
%>cooltools insulation --ignore-diags 2 --append-raw-scores --min-frac-valid-pixels 0.66 --min-dist-bad-bin 0 --chunksize 2000000
win= 100000
Creating GMS_Domains.bedpe: "hicMergeDomains" with all Strong boundaries of 4DN and "intersectBed" -a all_ArrowHeads -b Strong_boundaries
%>hicMergeDomains --value 5000
Creating GMS_Loops.bedgraph: "hicMergeLoops" of hicCups merged_loops + all_hicDetectLoops -r 25000
%>hicMergeLoops -r 25000
CTCF&RAD21
%>gimme scan -gc -f 0.1 -b -N 1
Boundaries
%>gimme motifs -b genomic -s 0 -f 0.2 -t Homer --known -p JASPAR2020_vertebrates -N 1
Loops
%>gimme motifs -b genomic -s 5000 -f 0.2 -t Homer --known -p JASPAR2020_vertebrates -N 1